- Home
- Services
- About
- News
- Contact
- Sound radix surfereq descarga foro x64
- How to find p value for hypothesis test calculator
- Sew what pro key
- Youtube marathi drama
- Permainan perang sniper
- Lee child jack reacher books
- Halis grotesque bold font
- Colnago c40 b stay
- Rabindranath ekhane kokhono khete asen ni watch online free
- Installing opengl 4-4 on mac
- Alpha zawgyi myanmar fonts for iphone
- Visual studio enterprise crack
- Siemens teamcenter for manufacturing
- Huey helicopter price
- Diabolik lovers game reiji route translation
- Pinnacle studio 16 ultimate updates
- Vrchat eye tracking
- How to laucnch quicken home inventory
- The sims 3 console commands
- Tame impala let it happen lyrics genius
- Ac3d plugin dds texture
- Diablo 2 lod cd k3ey
- X force autocad 2015
- The serum 2
- Watch badla movie online
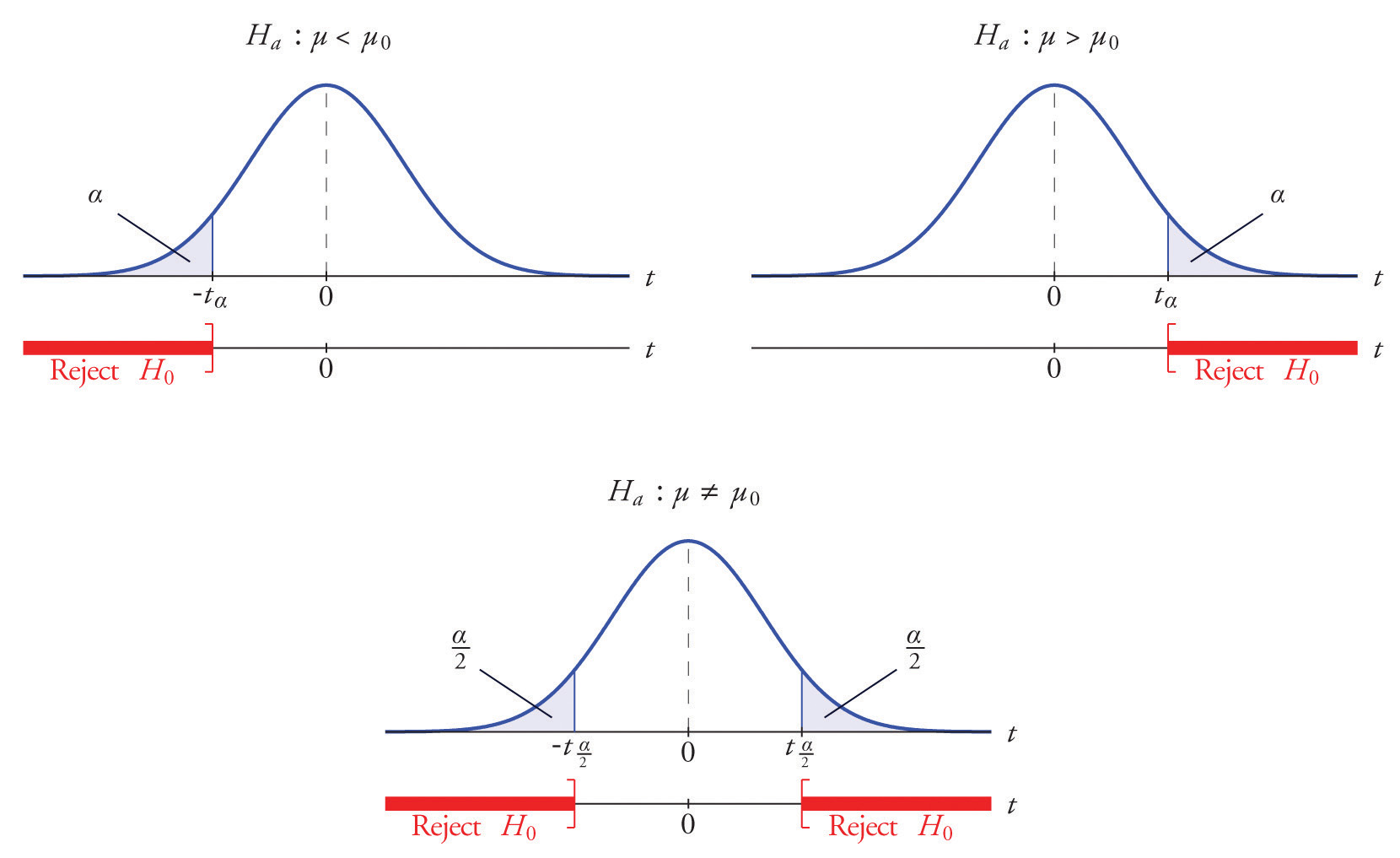

How to find p value for hypothesis test calculator free#
Another approach to the Monte Carlo estimation of significance would be to use the simulated test statistics to estimate the shape of the probability distribution and then to calculate a P value from this, but the use of ranks renders the process distribution free and is used almost universally. It is perhaps worth explicitly making the point that this procedure utilizes the ranks, rather than the actual values, of the test statistics. These realizations can be ranked, and then the probability, under the null hypothesis, that the test statistic from the actual data has the observed rank or a higher rank is ( r+1)/( n+1), the proportion of all possible rankings of the realizations that fulfill this criterion.

The reasoning is roughly as follows: if the null hypothesis is true, then the test statistics of the n replicates and the test statistic of the actual data are all realizations of the same random variable. However, Davison and Hinkley ( 1997) give the correct formula for obtaining an empirical P value as ( r+1)/( n+1). Typically, the estimate of the P value is obtained as, where n is the number of replicate samples that have been simulated and r is the number of these replicates that produce a test statistic greater than or equal to that calculated for the actual data. In this letter, we would first like to draw attention to the fact that some currently available genetic-analysis programs (including some of our own) use a method of calculating empirical P values that is not strictly correct. 1999), and a new test of linkage for a second locus conditional on information from an already-known locus (Cordell et al. Examples of procedures for genetic analysis that use simulation methods to determine statistical significance are CLUMP (Sham and Curtis 1995), MCETDT (Zhao et al. In contrast, Monte Carlo methods can be used to obtain an empirical P value that approximates the exact P value without relying on asymptotic distributional theory or exhaustive enumeration. The reasons for this include the following: (1) many test statistics do not have a standard asymptotic distribution (2) even if a standard asymptotic distribution does exist, it may not be reliable in realistic sample sizes and (3) calculation of the exact sampling distribution through exhaustive enumeration of all possible samples may be too computationally intensive to be feasible. It has become commonplace in the statistical analysis of genetic data to use Monte Carlo procedures to calculate empirical P values.
- Home
- Services
- About
- News
- Contact
- Sound radix surfereq descarga foro x64
- How to find p value for hypothesis test calculator
- Sew what pro key
- Youtube marathi drama
- Permainan perang sniper
- Lee child jack reacher books
- Halis grotesque bold font
- Colnago c40 b stay
- Rabindranath ekhane kokhono khete asen ni watch online free
- Installing opengl 4-4 on mac
- Alpha zawgyi myanmar fonts for iphone
- Visual studio enterprise crack
- Siemens teamcenter for manufacturing
- Huey helicopter price
- Diabolik lovers game reiji route translation
- Pinnacle studio 16 ultimate updates
- Vrchat eye tracking
- How to laucnch quicken home inventory
- The sims 3 console commands
- Tame impala let it happen lyrics genius
- Ac3d plugin dds texture
- Diablo 2 lod cd k3ey
- X force autocad 2015
- The serum 2
- Watch badla movie online
